A small step for this blog, but a big help for me…
After quite some time of inactivity (eg day-work) I revived this almost abandoned blog as test-case for playing with 2025 AI coding tools. My problem: as many blogs, the gaps between new articles became so long, that often needed to evolve the already minimal blog implementation. I realized that it took more time to upgrade hugo and adapt to the changes than the time I spent on writing actual content. ...
Hugo and org: experience some time after migration
I have now converted all my wordpress and drupal based blog sites to Hugo and Org format as input. Doom emacs has simplified this process as the standart config is already convenient to use and does only require minor tweeks (eg in case you would like to keep a few unpublished blog entries in a common org file).

Monty Hall problem - a small simulation in R
The Monty Hall problem is an interesting example for how much intuition can mislead us in some statistical contexts. Even more disturbing though is, for how long we are prepared to debate and defend an expected result before actually checking our initial guesses using a simple Monte Carlo simulation. Here is simple simulation implementation the Monty Hall game show problem: In the TV show “Let’s Make a Deal” the host Monty Hall would offer to game participant the choice of three doors. One of them was hiding a valuable price (eg a car) - behind the other two doors were only two less desirable goats. ...
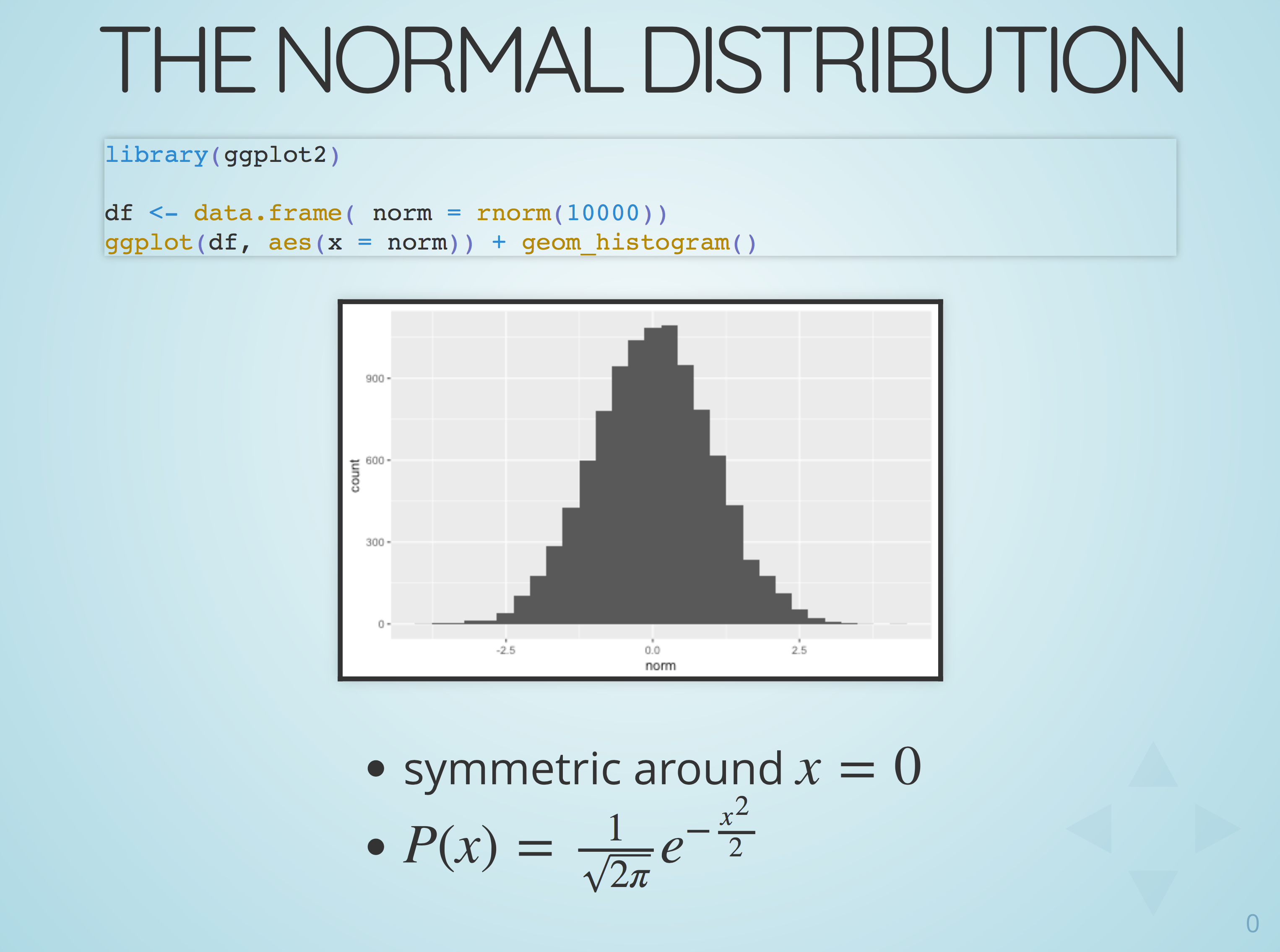
Slides and blog posts with R and emacs org-mode
Preparing a larger number of slides with R code and plots can be a bit tedious with standard desktop presentation software like powerpoint or keynote. The manual effort to change the example code, run the analysis and then cut and paste updated graphs, tables and code is high. Sooner or later one is bound to create inconsistencies between code and expected results or even syntax errors ...

Using Data Frames in Feather format (Apache Arrow)
Triggered by the RStudio blog article about feather I did the one line install and compared the results on a data frame of 19 million rows. First results look indeed promising: 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 # build the package > devtools::install_github("wesm/feather/R") # load an existing data frame (19 million rows with batch job execution results) > load("batch-12-2015.rda") # write it in feather format... > write_feather(dt,"batch-12-2015.feather") # ... which is not compressed, hence larger on disk > system("ls -lh batch-12-2015.*") -rw-r--r-- 1 dirkd staff 813M 7 Apr 11:35 batch-12-2015.feather -rw-r--r-- 1 dirkd staff 248M 27 Jan 22:42 batch-12-2015.rda # a few repeat reads on an older macbook with sdd > system.time(load("batch-12-2015.rda")) user system elapsed 8.984 0.332 9.331 > system.time(dt1 <- read_feather("batch-12-2015.feather")) user system elapsed 1.103 1.094 7.978 > system.time(load("batch-12-2015.rda")) user system elapsed 9.045 0.352 9.418 > system.time(dt1 <- read_feather("batch-12-2015.feather")) user system elapsed 1.110 0.658 3.997 > system.time(load("batch-12-2015.rda")) user system elapsed 9.009 0.356 9.393 > system.time(dt1 <- read_feather("batch-12-2015.feather")) user system elapsed 1.099 0.711 4.548 So, around half the elapsed time and about 1/10th of the user cpu time (uncompressed) ! Of course these measurements are from file system cache rather than the laptop SSD, but the reduction in wall time is nice for larger volume loads. ...

Setting up an RStudio server for iPad access
Sometimes it can be convenient to run RStudio remotely from an iPad or another machine with little RAM or disk space. This can be done quite easily using the free RStudio Server on OSX via docker. To do this: Find the rocker/rstudio image on docker hub and follow the setup steps here at github. Once the image is running, you should be able to connect with Safari on the host Mac to the login page eg at ...

Cached, asynchronous IP resolution
Resolving IP addresses to host names is quite helpful for getting a quick overview of who is connecting from where. This may need some care to not put too much strain on your DNS server with a large number of repeated lookups. Also you may not want to wait for timeouts on IPs that do not resolve. R itself is not supporting this specifically but can easily exploit asynchronous DNS lookup tools like adns (on OSX from homebrew) and provide a cache to speed things up. Here is a simple example for a vectorised lookup using a data.table as persistent cache. ...

Using R for weblog analysis
Apache Weblog Analysis Whether you run your own blog or web server or use some hosted service – at some point you may be interested in some information on how well your server or your users are doing. Many infos like hit frequency, geolocation of users and distribution of spent bandwidth are very useful for this and can be obtained in different ways: by instrumenting the page running inside the client browser (eg piwik) by analysis of the web server logs (eg webalizer) For the latter I have been using for several years webalizer, which does nice web based analysis plots. More recently I moved to a more complicated server environment with several virtual web services and I found the configuration and data selection options a bit limting. Hence I started as a toy project to implement the same functionality with a set of simple R scripts, which I will progressively share here. ...

Getting hold of remote weblogs
The last post was assuming that the weblogs to analyse are directly accessible by the R session which may not be the case if your analysis is running on a remote machine. Also in some cases you may want to filter out some uninteresting log records (eg local clients on the web server or local area accesses from known clients). The next examples show how to modify the previous R script using the R pipe function to take this into account: ...
Hadoop + kerberos with RStudio
While the setup from the previous posts works for the hadoop shell commands, you will still fail to access the remote cluster from GUI programs (eg RStudio) and/or with hadoop plugins like RHadoop. There are two reasons for that: GUI programs do not inherit your terminal/shell enviroment variables - unless you start them from a terminal session with $ open /Applications/RStudio.app $HADOOP_OPTS / $YARN_OPTS are not evaluated by other programs even if the variables are present in their execution environment. The first problem is well covered by various blog posts. The main difficulty is only to find the correct procedure for your OSX version,since Apple has changed several times over the years: ...
Connect to a remote, kerberized hadoop cluster
To use a remote hadoop cluster with kerberos authentication you will need to get a proper krb5.conf file (eg from your remote cluster /etc/kerb5.conf) and place the file /etc/krb5.conf on your client OSX machine. To use this configurations from your osx hadoop client change your .[z]profile to: export HADOOP_OPTS="-Djava.security.krb5.conf=/etc/krb5.conf" export YARN_OPTS="-Djava.security.krb5.conf=/etc/krb5.conf" With java 1.7 this should be sufficient to detect the default realm, the kdc and also any specific authentication options used by your site. Please make sure the kerberos configuration is already in place when you obtain your ticket with ...

Basic set-up of hadoop on OSX yosemite
Why would I do this? An OSX laptop will not allow to do any larger scale data processing, but it may be convenient place to develop/debug hadoop scripts before running on a real cluster. For this you likely want to have a local hadoop “cluster” to play with, and use the local commands as client for an larger remote hadoop cluster. This post covers the local install and basic testing. A second post shows how to extend the setup for accessing /processing against a remote kerberized cluster. ...